Some years in the past, when working as a marketing consultant, I used to be deriving a comparatively complicated ML algorithm, and was confronted with the problem of creating the interior workings of that algorithm clear to my stakeholders. That’s once I first got here to make use of parallel coordinates – as a result of visualizing the relationships between two, three, perhaps 4 or 5 variables is straightforward. However as quickly as you begin working with vectors of upper dimension (say, 13, for instance), the human thoughts oftentimes is unable to know this complexity. Enter parallel coordinates: a instrument so easy, but so efficient, that I usually surprise why it’s so little in use in on a regular basis EDA (my groups are an exception). Therefore, on this article, I’ll share with you the advantages of parallel coordinates primarily based on the Wine Dataset, highlighting how this method may help uncover correlations, patterns, or clusters within the knowledge with out shedding the semantics of options (e.g., in PCA).
What are Parallel Coordinates
Parallel coordinates are a standard methodology of visualizing high-dimensional datasets. And sure, that’s technically right, though this definition doesn’t absolutely seize the effectivity and magnificence of the tactic. In contrast to in a typical plot, the place you’ve two orthogonal axes (and therefore two dimensions which you can plot), in parallel coordinates, you’ve as many vertical axes as you’ve dimensions in your dataset. This implies an remark will be displayed as a line that crosses all axes at its corresponding worth. Need to study a elaborate phrase to impress on the subsequent hackathon? “Polyline”, that’s the right time period for it. And patterns then seem as bundles of polylines with related behaviour. Or, extra particularly: clusters seem as bundles, whereas correlations seem as trajectories with constant slopes throughout adjoining axes.
Surprise why not simply do PCA (Principal Element Evaluation)? In parallel coordinates, we retain all the unique options, which means we don’t condense the knowledge and challenge it right into a lower-dimensional house. So this eases interpretation rather a lot, each for you and to your stakeholders! However (sure, over all the thrill, there should nonetheless be a however…) you must take excellent care to not fall into the overplotting-trap. When you don’t put together the information rigorously, your parallel coordinates simply turn into unreadable – I’ll present you within the walkthrough that function choice, scaling, and transparency changes will be of nice assist.
Btw. I ought to point out Prof. Alfred Inselberg right here. I had the honour to dine with him in 2018 in Berlin. He’s the one who received me hooked on parallel coordinates. And he’s additionally the godfather of parallel coordinates, proving their worth in a mess of use circumstances within the Eighties.
Proving my Level with the Wine Dataset
For this demo, I selected the Wine Dataset. Why? First, I like wine. Second, I requested ChatGPT for a public dataset that’s related in construction to one among my firm’s datasets I’m at present engaged on (and I didn’t wish to tackle all the trouble to publish/anonymize/… firm knowledge). Third, this dataset is well-researched in lots of ML and Analytics functions. It accommodates knowledge from the evaluation of 178 wines grown by three grape cultivars in the identical area of Italy. Every remark has 13 steady attributes (assume alcohol, flavonoid focus, proline content material, color depth,…). And the goal variable is the category of the grape.
So that you can observe by, let me present you easy methods to load the dataset in Python.
import pandas as pd
# Load Wine dataset from UCI
uci_url = "https://archive.ics.uci.edu/ml/machine-learning-databases/wine/wine.knowledge"
# Outline column names primarily based on the wine.names file
col_names = [
"Class", "Alcohol", "Malic_Acid", "Ash", "Alcalinity_of_Ash", "Magnesium",
"Total_Phenols", "Flavanoids", "Nonflavanoid_Phenols", "Proanthocyanins",
"Color_Intensity", "Hue", "OD280/OD315", "Proline"
]
# Load the dataset
df = pd.read_csv(uci_url, header=None, names=col_names)
df.head()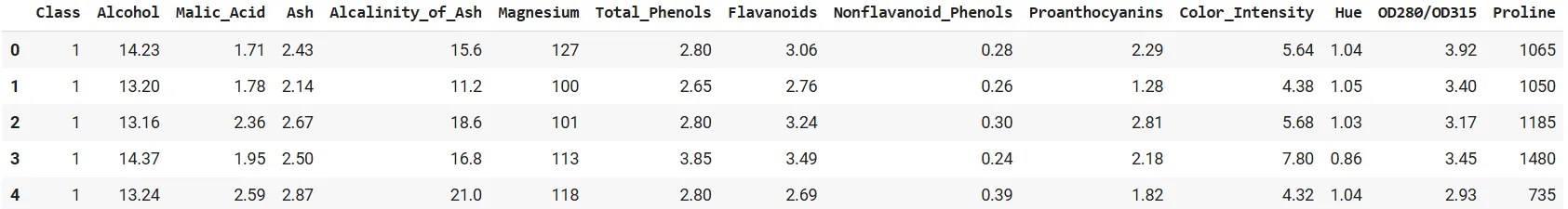
Good. Now, let’s derive a naïve plot as a baseline.
First Step: Constructed-In Pandas
Let’s use the built-in pandas plotting operate:
from pandas.plotting import parallel_coordinates
import matplotlib.pyplot as plt
plt.determine(figsize=(12,6))
parallel_coordinates(df, 'Class', colormap='viridis')
plt.title("Parallel Coordinates Plot of Wine Dataset (Unscaled)")
plt.xticks(rotation=45)
plt.present()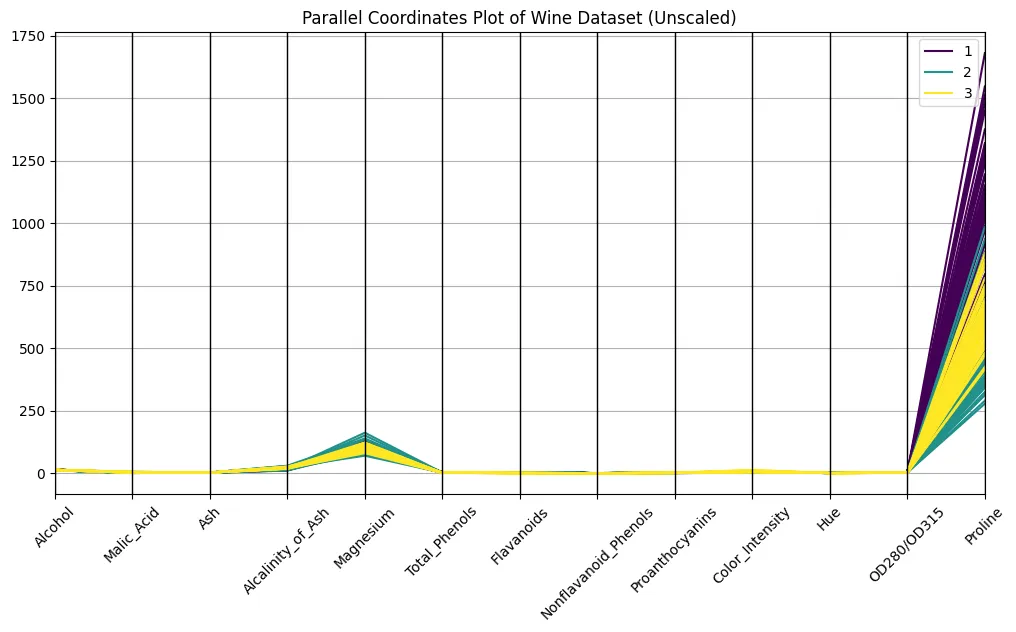
Seems to be good, proper?
No, it doesn’t. You definitely are in a position to discern the courses on the plot, however the variations in scaling make it onerous to check throughout axes. Examine the orders of magnitude of proline and hue, for instance: proline has a powerful optical dominance, simply due to scaling. An unscaled plot seems nearly meaningless, or no less than very troublesome to interpret. Even so, faint bundles over courses appear to look, so let’s take this as a promise for what’s but to return…
It’s all about Scale
A lot of you (everybody?) are conversant in the min-max scaling from ML preprocessing pipelines. So let’s not use that. I’ll do some standardization of the information, i.e., we do Z-scaling right here (every function could have a imply of zero and unit variance), to present all axes the identical weight.
from sklearn.preprocessing import StandardScaler
# Separate options and goal
options = df.drop("Class", axis=1)
scaler = StandardScaler()
scaled = scaler.fit_transform(options)
# Reconstruct a DataFrame with scaled options
scaled_df = pd.DataFrame(scaled, columns=options.columns)
scaled_df["Class"] = df["Class"]
plt.determine(figsize=(12,6))
parallel_coordinates(scaled_df, 'Class', colormap='plasma', alpha=0.5)
plt.title("Parallel Coordinates Plot of Wine Dataset (Scaled)")
plt.xticks(rotation=45)
plt.present()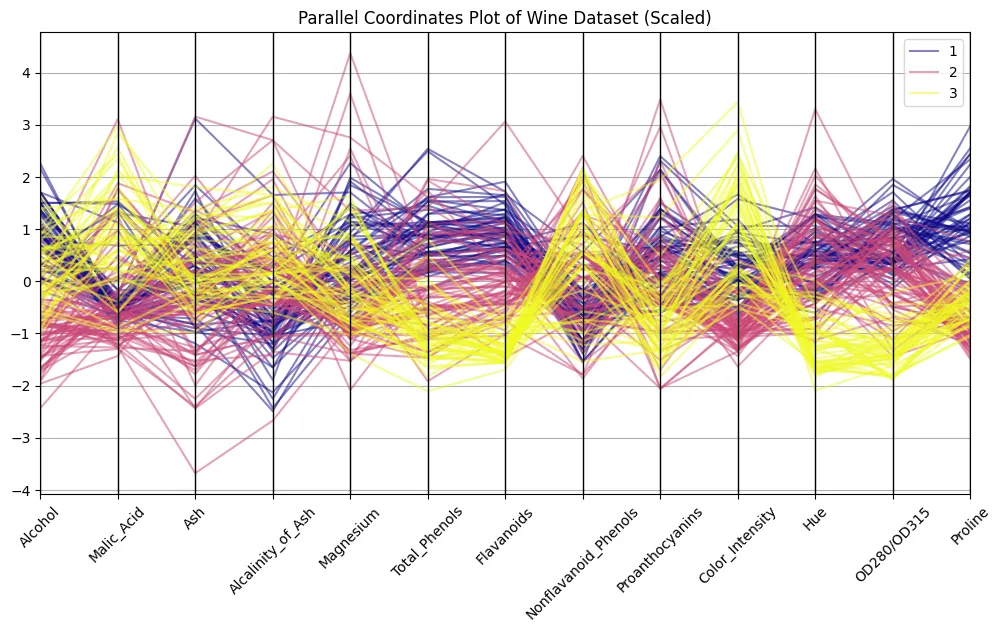
Bear in mind the image from above? The distinction is hanging, eh? Now we are able to discern patterns. Attempt to distinguish clusters of strains related to every wine class to search out out what options are most distinguishable.
Characteristic Choice
Did you uncover one thing? Right! I received the impression that alcohol, flavonoids, color depth, and proline present nearly textbook-style patterns. Let’s filter for these and attempt to see if a curation of options helps make our observations much more hanging.
chosen = ["Alcohol", "Flavanoids", "Color_Intensity", "Proline", "Class"]
plt.determine(figsize=(10,6))
parallel_coordinates(scaled_df[selected], 'Class', colormap='coolwarm', alpha=0.6)
plt.title("Parallel Coordinates Plot of Chosen Options")
plt.xticks(rotation=45)
plt.present()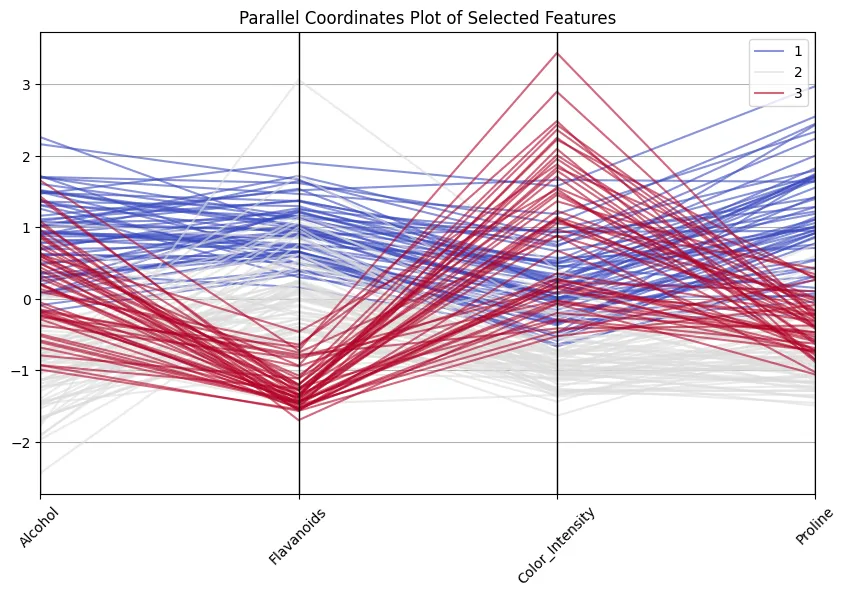
Good to see how class 1 wines all the time rating excessive on flavonoids and proline, whereas class 3 wines are decrease on these however excessive in color depth! And don’t assume that’s a useless train… 13 dimensions are nonetheless okay to deal with and to examine, however I’ve encountered circumstances with 100+ dimensions, making decreasing dimensions crucial.
Including Interplay
I admit: the examples above are fairly mechanistic. When writing the article, I additionally positioned hue subsequent to alcohol, which made my properly proven courses collapse; so I moved color depth subsequent to flavonoids, and that helped. However my goal right here was to not provide the excellent copy-paste piece of code; it was moderately to indicate you using parallel coordinates primarily based on some easy examples. In actual life, I’d arrange a extra explorative frontend. Plotly parallel coordinates, for example, include a “brushing” function: there you may choose a subsection of an axis and all polylines falling inside that subset will probably be highlighted.
You may as well reorder axes by easy drag and drop, which frequently helps reveal correlations that had been hidden within the default order. Trace: Strive adjoining axes that you just suspect to co-vary.
And even higher: scaling isn’t vital for inspecting the information with plotly: the axes are mechanically scaled to the min- and max values of every dimension.
Right here’s a code so that you can reproduce in your Colab:
import plotly.categorical as px
# Preserve class as a separate column; Plotly's parcoords expects numeric color for 'colour'
df["Class"] = df["Class"].astype(int)
fig_all = px.parallel_coordinates(
df,
colour="Class", # numeric color mapping (1..3)
dimensions=options.columns,
labels={c: c.substitute("_", " ") for c in scaled_df.columns},
)
fig_all.update_layout(
title="Interactive Parallel Coordinates — All 13 Options"
)
# The file under will be opened in any browser or embedded by way of <iframe>.
fig_all.write_html("parallel_coordinates_all_features.html",
include_plotlyjs="cdn", full_html=True)
print("Saved:")
print(" - parallel_coordinates_all_features.html")
# present figures inline
fig_all.present()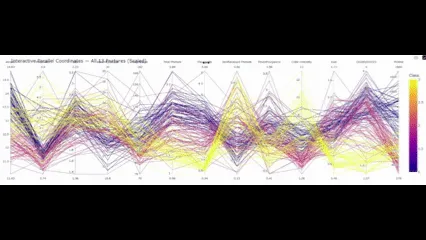
So with this closing aspect in place, what conclusions will we draw?
Conclusion
Parallel coordinates are usually not a lot in regards to the onerous numbers, however far more in regards to the patterns that emerge from these numbers. Within the Wine dataset, you might observe a number of such patterns – with out working correlations, doing PCA, or scatter matrices. Flavonoids strongly assist distinguish class 1 from the others. Color depth and hue separate courses 2 and three. Proline additional reinforces that. What follows from there may be not solely which you can visually separate these courses, but in addition that it offers you an intuitive understanding of what separates cultivars in follow.
And that is precisely the energy over t-SNE, PCA, and so forth., these methods challenge knowledge into elements which can be glorious in distinguishing the courses… However good luck attempting to elucidate to a chemist what “part one” means to him.
Don’t get me fallacious: parallel coordinates are usually not the Swiss military knife of EDA. You want stakeholders with an excellent grasp of knowledge to have the ability to use parallel coordinates to speak with them (else proceed utilizing boxplots and bar charts!). However for you (and me) as an information scientist, parallel coordinates are the microscope you’ve all the time been eager for.
Steadily Requested Questions
A. Parallel coordinates are primarily used for exploratory evaluation of high-dimensional datasets. They help you spot clusters, correlations, and outliers whereas preserving the unique variables interpretable.
A. With out scaling, options with giant numeric ranges dominate the plot. Standardising every function to imply zero and unit variance ensures that each axis contributes equally to the visible sample.
A. PCA and t-SNE scale back dimensionality, however the axes lose their authentic which means. Parallel coordinates preserve the semantic hyperlink to the variables, at the price of some muddle and potential overplotting.
Login to proceed studying and revel in expert-curated content material.

